 Michael P. Sheetz, PhD
Michael P. Sheetz, PhD
Robert A. Welch Distinguished Chair in Chemistry
Professor, Department of Biochemistry & Molecular Biology
Tel: (409) 747-0452
E-mail: misheetz@utmb.edu
Mail Route: 0645
Research
The morphology of cells, organs and whole organisms is determined by the generation of forces on the immediate environment, which is either extracellular matrix or adjacent cells. We are currently engaged in studies to understand the detailed molecular mechanisms involved in a variety of. cellular phenomena from cancer metastasis to aging and muscle function. Recent studies defined the major components of the matrix rigidity sensor, which is missing in all tumor cells and makes them transformed. The state change with transformation has revealed new ways to treat cancer selectively that are under implementation. Senescent cells also have dramatic changes in the way that they respond to external forces that may unlock new ways to rejuvenate or remove senescent cells. Mechanics provides new ways to modify cell behavior without drugs and may enable treatment of many disorders including cancer.
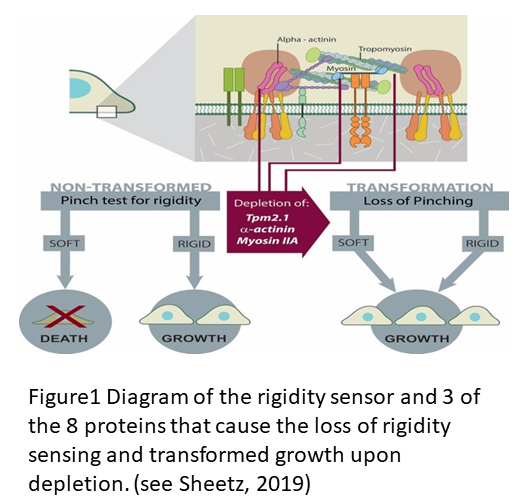
The description of the process of rigidity sensing has been reviewed recently (see Sheetz, 2019 and Wolfenson et al., 2019) but there is a lot more to learn about the process that is fundamental to cell differentiation and cancer cell transformation. The diagram in Figure 1 shows the important elements in the two micrometer long sarcomere like complex that senses rigidity in virtually all cell types that have been tested except for cancer cells. Major mysteries remain about the detailed process that causes assembly of the contractile apparatus, limits the contraction to ~100 nm and causes the relaxation and disassembly all within a minute as illustrated https://www.youtube.com/watch?v=xAruFbqU7Wg&feature=youtu.be
Surprisingly, in the rigidity sensing units, very large forces are generated by the nonmuscle myosin that can be more than ten-fold greater than myosin stall forces measured in vitro. The mechanism of contraction involves a stochastic accumulation of the myosin heads in a high energy state on actin that then results in a 2.5 nm movement. This treatment may be particularly relevant to muscle myosin contractions under isometric loads. (https://mbi.nus.edu.sg/science-features/myosin-motors-show-their-muscle/)
Our recent studies show that integrin clusters can form on thin matrix fibers if the fibers are crossing or closely aligned. The clusters are the same size as on large 2D surfaces and this further reinforces the concept that the basic 100 nm integrin clusters are the basis of adhesions both in 3D environments as well as 2D. Further, the majority of the integrins are not bound to ligand. These studies form the basis of further analyses of the mechanism of adhesion assembly and dynamics. (https://mbi.nus.edu.sg/science-features/when-spacing-matters/)
The repair of DNA damage appears to involve the rapid recruitment of inositol lipids to the site of damage in a fraction of a second. This is an important step in ATR mediated DNA damage repair and may link to other damage systems involving p53. Surprisingly, there are no membranes in these regions of the nucleus and the lipids appear to be carried by steroid binding proteins that have hydrophobic pockets. (https://mbi.nus.edu.sg/science-features/lipid-mediated-dna-repair/)
